Prologue
Prologue
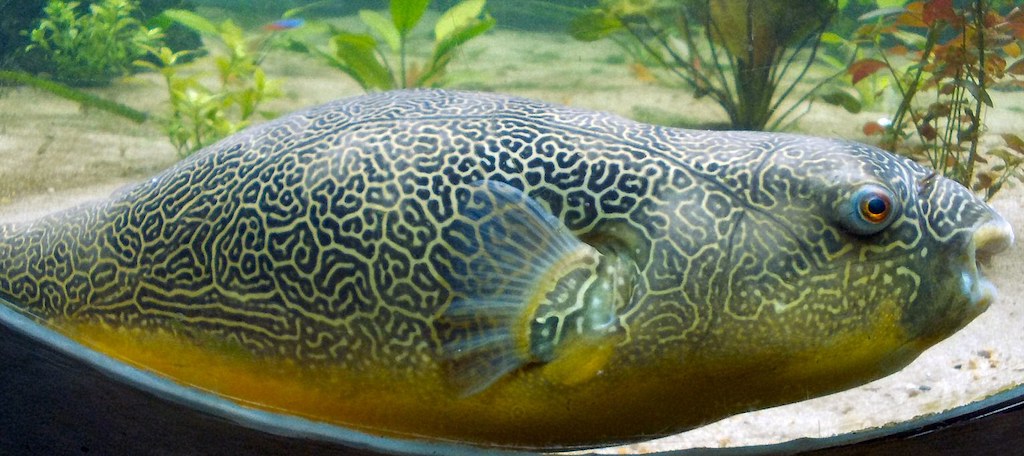
Random Walks and Turing Patterns
Why do zebras have stripes? Simple molecular interactions give rise to reaction-diffusion systems that spontaneously produce the complex, self-organized patterns found across the biological world.
Begin the course
 Module 1
Module 1
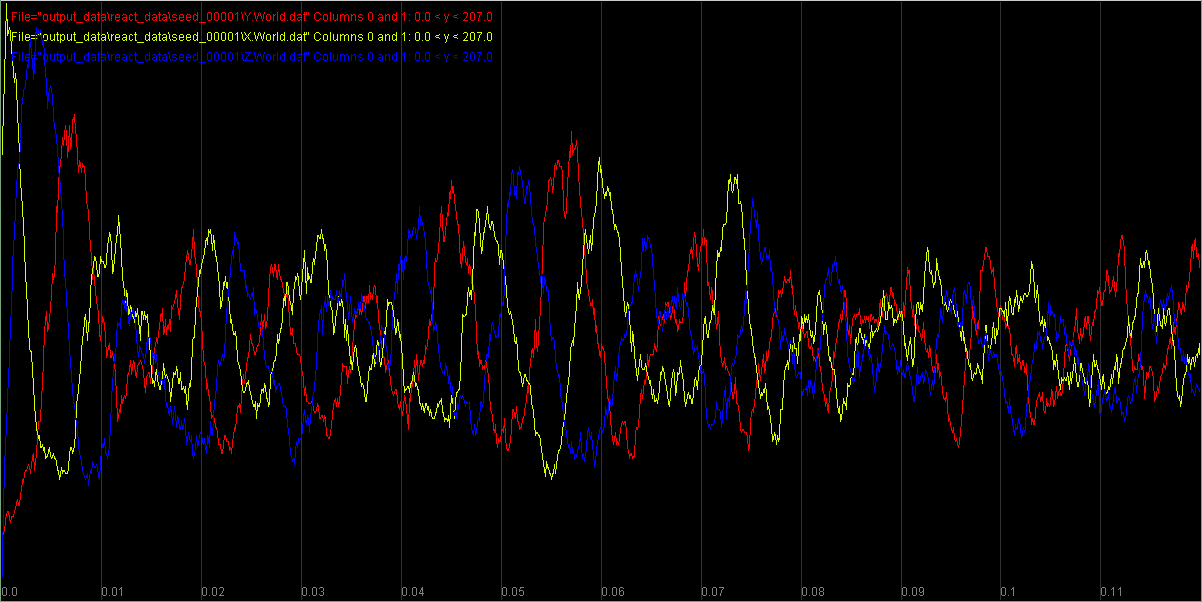
Finding Motifs in Transcription Factor Networks
Transcription factor networks exhibit recurring structural motifs, including oscillators, that appear too often to be coincidental. We'll model these patterns and ask why evolution keeps arriving at the same solutions.
Read more
 Module 2
Module 2
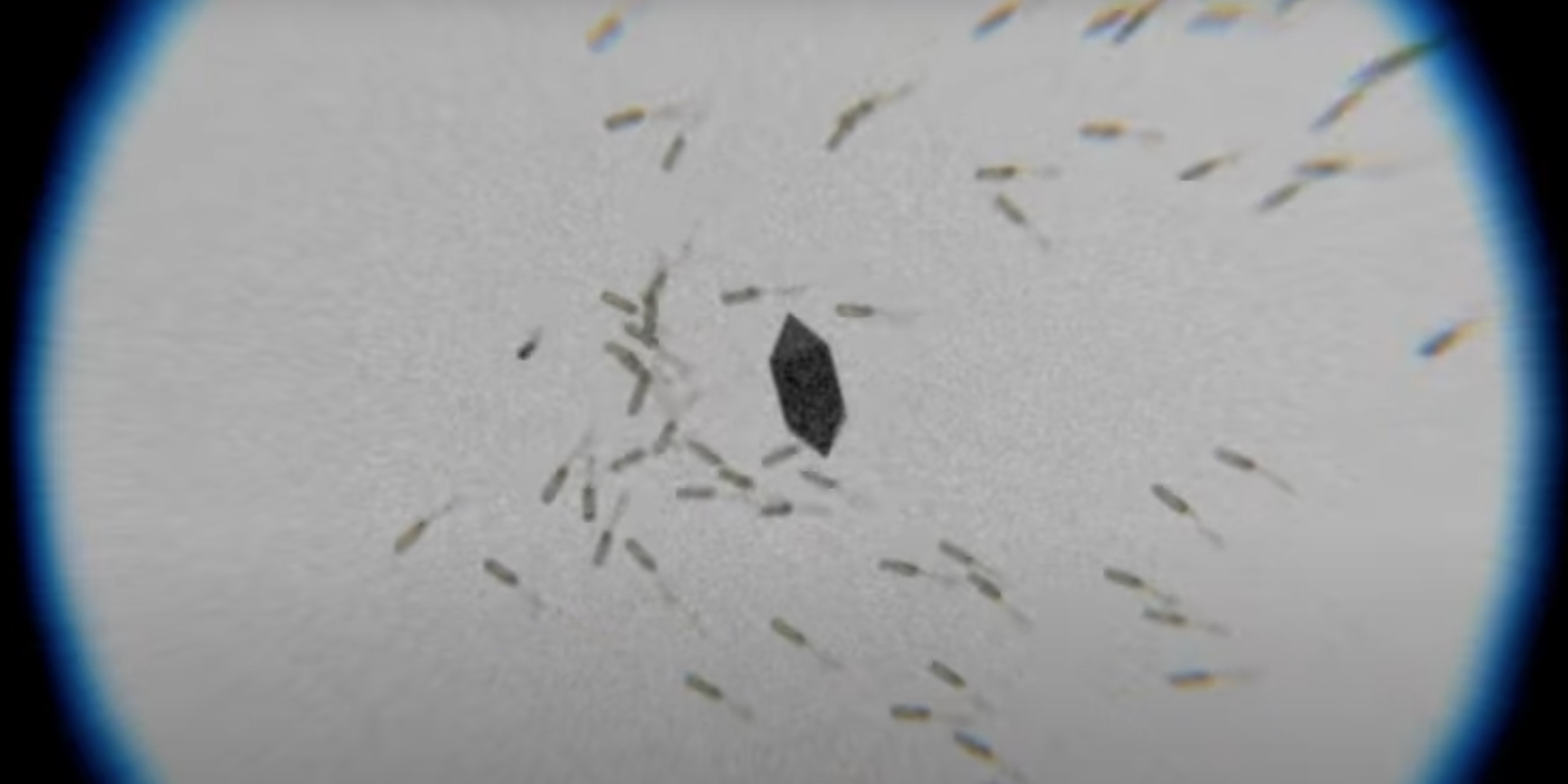
Unpacking E. coli's Genius Exploration Algorithm
Bacteria navigate chemical gradients through a cascade of molecular reactions that produce robust, adaptive behavior. We'll construct a model of this system, introduce perturbations, and examine how the cell maintains its strategy.
Read more

Each module in Biological Modeling is built around a real question in computational biology, and we use real tools to solve it. Bringing this work to the world with a student team has been one of the great joys of my career.
A Philomath Course
Stay connected
Join thousands of learners across our growing library of free courses.